 This hypothesis is supported by the differentiation-promoting effects of CDF4 overexpression and implies that wild-type CDF4 primarily promotes the exit of the distal daughter cells from the stem cell state.In eukaryotes, DNA is packaged into chromatin with the core histone proteins including histone H2A, H2B, H3, and H4. 
We therefore consider it a more straightforward interpretation that VP16-CDF4 expression slows down the differentiation of the distal stem cell daughters. However, the fact that the absence of CDF4 is essential to maintain the integrity of the mitotically inactive QC is at odds with this interpretation. First, the division rate of the CSCs could be increased, implying that native CDF4 would primarily repress cell division. The increase of two stem-cell-like layers in roots expressing VP16-CDF4 can be explained in two ways. In the wild-type root, the occasional presence of two CSC-like layers plausibly represents a transient stage immediately after stem cell division before the distal daughter cell destined to produce columella cells has differentiated. We hypothesize the latter effect to be more pronounced in complete CDF4 loss-of-function roots, because in the wild-type background, VP16-ΔNCDF4 protein must compete with endogenous CDF4. What is the precise role of CDF4 in the columella? Misexpression experiments indicate that CDF4 promotes columella cell differentiation, and by antagonizing CDF4 repressor activity, we find a significant increase of roots with double stem-cell-like layers. In summary, we conclude that TPL/TPRs and HDA19 are required for the direct repression of CDF4 through WOX5 in the root stem cell niche. Importantly, tpl-1 and hda19-3 alleviate the transcriptional repression of pCDF4:3xnlsGFP by WOX5 overexpression ( Figure 7E). This is not the case for the hda19-3 mutant, which, however, displays strongly reduced pCDF4:3xnlsGFP expression throughout the root meristem ( Figures 7C and 7D), consistent with HDA19 requirement in additional developmental processes. Additionally, the expression of pCDF4:3xnlsGFP is significantly upregulated at the CSC position in tpl-1 ( Figure 7D). The low penetrance of this effect in hda19-3 might be due to genetic redundancy, which is evident, for example, from the embryo lethal phenotype of hda19-3 hda6/rts1 double mutants (data not shown). Similar to the situation in wox5-1, we find that pCDF4:3xnlsGFP is ectopically expressed in the QC position of tpl-1 and in hda19-3 mutants ( Figures 7A–7C). Next we tested whether TPL/TPRs and HDA19 are required for the direct repression of CDF4 by WOX5. The resulting proximal-distal gradient of WOX5-GFP protein with highest levels in the QC, low levels in the CSCs, and no detectable expression in the differentiated columella cells is reversely oriented to the observed CDF4 reporter expression gradient. Thus, we conclude that WOX5-GFP protein moves from the QC into the CSCs where it represses CDF4 expression, as indicated by the upregulation of the pCDF4mBS2:3xnlsGFP reporter in wild-type roots ( Figures 3F and 3H). 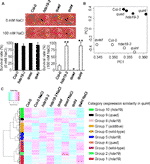
Importantly, we detect WOX5-GFP protein accumulation not only in the QC but also in the nuclei of the CSCs ( Figure 4B), whereas expression of the transcriptional control is in the QC but absent from the CSCs ( Figure 4A). First, we compared the signals of a transcriptional ( pWOX5:erCFP) and a translational ( pWOX5:WOX5-GFP) reporter using the WOX5 promoter. Complementation of the wox5-1 CSC defect was judged by the reappearance of a starch-free layer below the QC and a wild-type-like layered organization of the columella. To this end, we studied localization of WOX5 fused to green or yellow florescence protein (GFP, YFP), both of which fully complement the wox5-1 mutant when expressed from the endogenous WOX5 promoter ( Figures 4E–4G and 4L), indicating that the fusion proteins are functional. ), we wondered how CDF4 repression occurs there. Our results show that chromatin-mediated repression of differentiation programs is a common strategy in plant and animal stem cell niches. We further show that WOX5 represses CDF4 transcription by recruiting TPL/TPR co-repressors and the histone deacetylase HDA19, which consequently induces histone deacetylation at the CDF4 regulatory region. This creates a gradient of CDF4 transcription, which promotes differentiation opposite to the WOX5 gradient, allowing stem cell daughter cells to exit the stem cell state. 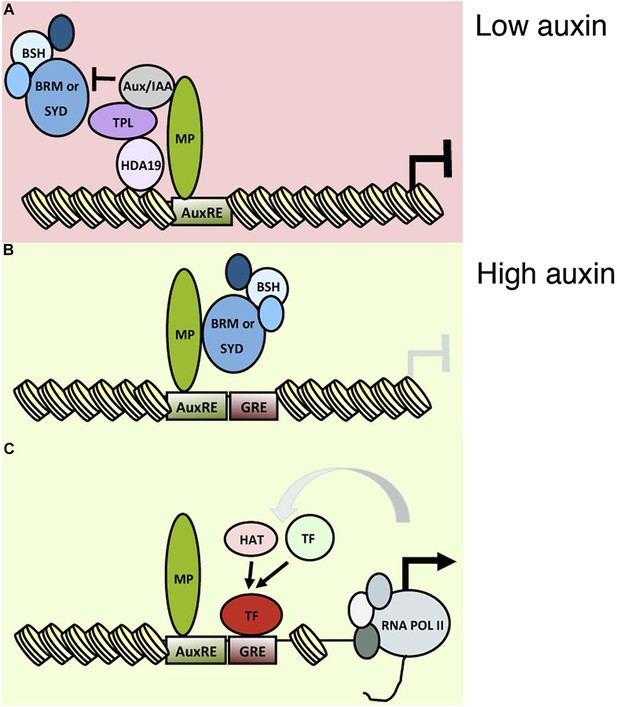
Here we show that in the Arabidopsis root meristem, the WOX5 protein moves from the root niche organizer, the quiescent center, into the columella stem cells, where it directly represses the transcription factor gene CDF4. In plants, WUSCHEL HOMEOBOX (WOX) transcription factors are central regulators of stem cell maintenance in different meristem types, yet their molecular mode of action has remained elusive. 
Stem cells in plants and animals are maintained pluripotent by signals from adjacent niche cells. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
